Khác biệt giữa bản sửa đổi của “DNA polymerase”
Không có tóm lược sửa đổi |
Không có tóm lược sửa đổi |
||
| Dòng 1: | Dòng 1: | ||
{{infobox enzyme |
|||
[[Tập tin:DNA polymerase.png|nhỏ|200px|DNA polymerase 3D structure.]] |
|||
| Name = DNA-directed DNA polymerase |
|||
'''DNA polymerase''' ('''ADN polymeraza''') là một enzyme tham gia chính vào quá trình [[nhân đôi DNA]]. Những enzyme này xúc tác cho quá trình polymer hóa các [[deoxiribonucleotide]] dựa trên trình tự của một chuỗi [[ADN|DNA]] khác. Sợi DNA mới được tạo thành sẽ bổ sung với [[sợi DNA khuôn]] theo [[nguyên tắc bổ sung]]. |
|||
| EC_number = 2.7.7.7 |
|||
| CAS_number = 9012-90-2 |
|||
| IUBMB_EC_number = 2/7/7/7 |
|||
| GO_code = 0034061 |
|||
| image = DNA polymerase.png |
|||
| width = 260px |
|||
| caption = 3D structure of the DNA-binding [[helix-turn-helix]] motifs in human DNA polymerase beta (based on PDB file [http://www.rcsb.org/pdb/explore.do?structureId=7ICG 7ICG]) |
|||
}} |
|||
The '''DNA polymerases''' are [[enzyme]]s that create [[DNA]] [[molecule]]s by assembling [[nucleotides]], the building blocks of DNA. These enzymes are essential to [[DNA replication]] and usually work in pairs to create two identical DNA strands from a single original DNA molecule. During this process, DNA polymerase “reads” the existing DNA strands to create two new strands that match the existing ones. <ref>{{cite journal | title = Calf thymus polymerase |author = Bollum, F.J. |journal = J. Biol. Chem. |year = 1960 |volume = 235 |pages = 2399–2403 |pmid = 13802334}}</ref><ref>{{cite journal | title = Biochemical studies of bacterial sporulation. II. Deoxy-ribonucleic acid polymerase in spores of ''Bacillus subtilis'' |author = Falaschi, A. and Kornberg, A. |journal = J. Biol. Chem. |year = 1966 |volume = 241 |pages = 1478–1482 |pmid = 4957767}}</ref><ref>{{cite journal | title = Enzymatic synthesis of deoxyribonucleic acid. I. Preparation of substrates and partial purification of an enzyme from ''Escherichia coli'' |author = Lehman, I.R., Bessman, M.J., Simms, E.S. and Kornberg, A. |journal = J. Biol. Chem. |year = 1958 |volume = 233 |pages = 163–170 |pmid = 13563462}}</ref><ref>{{cite journal | title = Enzymatic synthesis of deoxyribonucleic acid. XIV. Further purification and properties of deoxyribonucleic acid polymerase of ''Escherichia coli'' |author = Richardson, C.C., Schildkraut, C.L., Aposhian, H.V. and Kornberg, A. |journal = J. Biol. Chem. |year = 1964 |volume = 239 |pages = 222–232 |pmid = 14114848}}</ref><ref>{{cite journal | title = Enzymatic synthesis of deoxyribonucleic acid. VII. Synthesis of a polymer of deoxyadenylate and deoxythymidylate |author = Schachman, H.K., Adler, J., Radding, C.M., Lehman, I.R. and Kornberg, A. |journal = J. Biol. Chem. |year = 1960 |volume = 235 |pages = 3242–3249 |pmid = 13747134}}</ref><ref>{{cite journal | title = Purification and properties of deoxyribonucleic acid polymerase from ''Micrococcus lysodeikticus'' |author = Zimmerman, B.K. |journal = J. Biol. Chem. |year = 1966 |volume = 241 |pages = 2035–2041 |pmid = 5946628}}</ref> |
|||
This enzyme [[catalysis|catalyses]] the following [[chemical reaction]] |
|||
Tất cả DNA polymerase đều tổng hợp DNA theo hướng từ đầu 5' đến đầu 3' của DNA (5' và 3' là vị trí của nguyên tử [[cacbon|các bon]]). Cho tới nay, không có enzyme DNA polymerase nào là có khả năng tổng hợp một chuỗi mới (''de novo'') mà không cần [[hiđrôxít|nhóm OH]] ở đầu 3' có sẵn (tính chất này chỉ có ở những [[RNA polymerase]]). Vì lý do này mà DNA polymerase cần đến sự có mặt của các [[primer]], từ đó enzyme DNA polymerase có thể gắn các nucleotid tiếp theo. Các primer chính là các đoạn [[oligonucleic]] gồm các [[bazơ]] của [[ARN|RNA]] và [[ADN|DNA]] trong đó hai bazơ đầu tiên luôn là bazơ của RNA. Các primer này được tổng hợp nhờ hoạt động của một enzyme khác gọi là [[primase]]. Enzyme [[helicase]] cần thiết cho quá trình tháo xoắn của cấu trúc xoắn kép DNA để trở thành những sợi đơn giúp cho quá trình sao chép DNA diễn ra theo đúng [[nguyên tắc bán bảo toàn]] (''semiconservative'') |
|||
: [[deoxynucleoside]] triphosphate + DNA<sub>n</sub> <math>\rightleftharpoons</math> [[diphosphate]] + DNA<sub>n+1</sub> |
|||
Các DNA polymerase thường có tính bảo thủ cao về mặt cấu trúc, nghĩa là cấu trúc bộ phận tham gia xúc tác của enzyme này nhìn chung thay đổi rất ít giữa các loài. Những cấu trúc được bảo lưu này nói lên đặc điểm ưu việt trong quá trình [[tiến hóa]]. |
|||
Catalyses DNA-template-directed extension of the 3'- end of a DNA strand by one nucleotide at a time. |
|||
DNA polymerase được xem như là [[holoenzyme]] vì nó cần có một ion [[Mg]] làm vai trò một [[co-factor]] giúp cho hoạt động được chuẩn xác. Khi không có nguyên tử Mg này phần còn lại của enzyme gọi là một [[apoenzyme]]. |
|||
Every time a [[Cell division|cell divides]], DNA polymerase is required to help duplicate the cell’s DNA, so that a copy of the original DNA molecule can be passed to each of the daughter cells. In this way, genetic information is transmitted from generation to generation. |
|||
Chỉ có một vài DNA polymerase có khả năng [[Sửa chữa DNA|sửa chữa]] những sai hỏng trên sợi DNA mới tổng hợp trong quá trình nhân đôi. Khi phát hiện một cặp bazơ liên kết không đúng theo [[nguyên tắc bổ sung]], DNA polymerase sẽ trượt ngược trở lại và thực hiện hoạt tính [[exonuclease]] để cắt bỏ bazơ ngoài cũng của chuỗi [[axit nucleic|nucleic acid]] theo hướng 3'-> 5'cho đến vị trí của cặp bazơ sai hỏng (hoạt động này được gọi là [[hoạt tính đọc sửa]] (''proofreading'')). Sau khi cắt bỏ cặp base sai, enzyme polymerase sẽ thay thế bằng bazơ thích hợp và qua trình sao chép lại được tiến hành. |
|||
Before replication can take place, an enzyme called [[helicase]] unwinds the DNA molecule from its tightly woven form. This opens up or “unzips” the double-stranded DNA to give two single strands of DNA that can be used as templates for replication. |
|||
Một vài loại [[virus]] cũng mã hóa những phân tử DNA polymerase đặc biệt có khả năng sao chép chọn lọc DNA của virus thông qua nhiều cơ chế khác nhau. Các [[retrovirus]] mã hóa một loại DNA polymerase sử dụng khuôn là RNA để tổng hợp những phân tử DNA, đó là các [[phiên mã ngược|enzyme phiên mã ngược]] (''reverse transcriptase''). |
|||
== |
==History== |
||
In 1956, [[Arthur Kornberg]] and colleagues discovered the enzyme [[DNA polymerase I]], also known as Pol I, in ''[[Escherichia coli]]''. They described the DNA replication process by which DNA polymerase copies the base sequence of a template DNA strand. Subsequently, in 1959, Kornberg was awarded the [[Nobel Prize in Physiology or Medicine]] for this work.<ref>{{cite web | title=The Nobel Prize in Physiology or Medicine 1959 | url=http://www.nobelprize.org/nobel_prizes/medicine/laureates/1959/ | publisher=[[Nobel Foundation]] | accessdate=December 1, 2012}}</ref> [[DNA polymerase II]] was also discovered by Kornberg and Malcolm E. Gefter in 1970 while further elucidating the role of Pol I in ''E. coli'' DNA replication.<ref name="pmid7908652">{{cite journal | author = Tessman I, Kennedy MA | title = DNA polymerase II of Escherichia coli in the bypass of abasic sites in vivo | journal = Genetics | volume = 136 | issue = 2 | pages = 439–48 |date=February 1994 | pmid = 7908652 | pmc = 1205799 | doi = }}</ref> |
|||
[[Vi khuẩn]] có ba nhóm DNA polymerase: |
|||
* '''[[DNA polymerase I|Pol I]]''': Đóng vai trò trong sửa chữa DNA. |
|||
* '''[[Pol II]]''': Hoạt động như một 5'->3' exonuclease (có thể loại bỏ primer), sau đó tiếp tục kéo dài phân tử DNA để lấp vào chỗ khuyết đó. |
|||
* '''[[Pol III]]''': Là polymerase chủ yếu của vi khuẩn (tham gia nhân đôi phân tử DNA), và có hoạt tính đọc sửa (''proofreading''). |
|||
== |
==Function== |
||
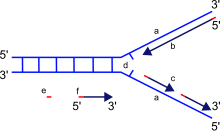
[[File:Replication fork.svg|thumb|DNA polymerase moves along the old strand in the 3'-5' direction, creating a new strand having a 5'-3' direction.]] |
|||
Cho đến bây giờ người ta đã phát hiện có 6 nhóm DNA Polymerase chính trong tế bào [[sinh vật nhân chuẩn|eukaryote]]: |
|||
[[File:DNA polymerase.svg|thumb|200px|right|DNA polymerase with proofreading ability]] |
|||
* '''RNA polimerase''': có vai trò như primase (để tổng hợp RNA primer), và sau đó thực hiện việc kéo dại phân tử DNA từ primer bằng cách gắn các nucleotid vào. Sau đó một vài trăm nucleotid kéo dài đó được cắt bỏ bởi Pol δ và ε |
|||
The main function of DNA polymerase is to make DNA from nucleotides, the building blocks of DNA. The DNA copies are created by the pairing of nucleotides to bases present on each strand of the original DNA molecule. This pairing always occurs in specific combinations, with [[cytosine]] along with [[guanine]], and [[thymine]] along with [[adenine]], forming two separate pairs, respectively. |
|||
* '''Pol III''': là enzim quan trọng nhất trong quá trình nhân đôi DNA, chức năng tổng hợp mạch DNA con bằng cách bổ sung dNTP vào đoạn mồi --> kéo dài đoạn mồi ra. Chúng là enzim hoạt động ở cả 2 mạch DNA. |
|||
* '''DNA polymerase I''': ở eukaryote có thể cắt bỏ primers (hoạt tính 5'->3' exonuclease), sau đó liền tổng hợp bù 1 đoạn DNA lấp vào khoảng trống đó. Ngoài ra DNA pol I còn tham gia hoạt tính sửa sai tích cực nhờ hoạt tính exonuclease 5' - 3'. |
|||
* '''Pol II, IV, V''': vai trò sửa sai trong nhân đôi DNA... |
|||
When creating DNA, DNA polymerase can add free nucleotides only to the 3' end of the newly forming strand. This results in elongation of the newly forming strand in a [[5'-3']] direction. No known DNA polymerase is able to begin a new chain (''de novo''); it can only add a nucleotide onto a pre-existing 3'-[[hydroxide|OH group]], and therefore needs a [[primer (molecular biology)|primer]] at which it can add the first nucleotide. Primers consist of [[RNA]] or DNA bases (or both). In DNA replication, the first two bases are always RNA, and are synthesized by another enzyme called [[primase]]. Enzymes,[[helicase]] and [[topoisomerase II]], are required to unwind DNA from a double-strand structure to a single-strand structure to facilitate replication of each strand consistent with the [[semiconservative]] model of DNA replication. |
|||
== Xem thêm == |
|||
* [[PCR]] |
|||
It is important to note that the [[directionality (molecular biology)|directionality]] of the newly forming strand (the daughter strand) is opposite to the direction in which DNA polymerase moves along the template strand. Since DNA polymerase requires a free 3' OH group for initiation of synthesis, it can synthesize in only one direction by extending the 3' end of the preexisting nucleotide chain. Hence, DNA polymerase moves along the template strand in a 3'-5' direction, and the daughter strand is formed in a 5'-3' direction. This difference enables the resultant double-strand DNA formed to be composed of two DNA strands that are [[antiparallel (biochemistry)|antiparallel]] to each other. |
|||
==Tham khảo== |
|||
{{tham khảo}} |
|||
== Liên kết ngoài == |
|||
* [http://www.jbc.org/cgi/content/full/276/47/43487 Eukaryotic DNA Polymerases: Proposal for a Revised Nomenclature] |
|||
* [http://arjournals.annualreviews.org/doi/abs/10.1146/annurev.biochem.71.090501.150041 Annual Review of Biochemistry: EUKARYOTIC DNA POLYMERASES] |
|||
* [http://nist.rcsb.org/pdb/molecules/pdb3_1.html DNA polymerase: PDB molecule of the month] |
|||
The function of DNA polymerase is not quite perfect, with the enzyme making about one mistake for every billion base pairs copied. Error correction is a property of some, but not all, DNA polymerases. This process corrects mistakes in newly synthesized DNA. When an incorrect base pair is recognized, DNA polymerase moves backwards by one base pair of DNA. The 3'-5' [[exonuclease]] activity of the enzyme allows the incorrect base pair to be excised (this activity is known as ''[[proofreading (biology)|proofreading]]''). Following base excision, the polymerase can re-insert the correct base and replication can continue forwards. This preserves the integrity of the original DNA strand that is passed onto the daughter cells. |
|||
{{Sinh học}} |
|||
===Structure=== |
|||
[[Thể loại:EC 2.7.7]] |
|||
The known DNA polymerases have highly conserved structure, which means that their overall catalytic [[Protein subunit|subunit]]s vary very little from species to species, independent of their domain structures. Conserved structures usually indicate important, irreplaceable functions of the cell, the maintenance of which provides evolutionary advantages. The shape can be described as resembling a right hand with thumb, finger, and palm domains. The palm domain appears to function in catalyzing the transfer of [[phosphoryl|phosphoryl group]]<nowiki/>s in the phosphoryl transfer reaction. DNA is bound to the palm when the enzyme is active. This reaction is believed to be catalyzed by a two-metal-ion mechanism. The finger domain functions to bind the [[Nucleotide triphosphates|nucleotide triphosphate]] with the template base. The thumb domain plays a potential role in the processivity, translocation, and positioning of the DNA.<ref name="pmid10364165">{{cite journal | author = Steitz TA | title = DNA polymerases: structural diversity and common mechanisms | journal = J. Biol. Chem. | volume = 274 | issue = 25 | pages = 17395–8 |date=June 1999 | pmid = 10364165 | doi = 10.1074/jbc.274.25.17395}}</ref> |
|||
[[Thể loại:Di truyền học]] |
|||
[[Thể loại:Enzyme]] |
|||
===Processivity=== |
|||
[[Thể loại:ADN]] |
|||
DNA polymerase’s rapid catalysis is due to its processive nature. [[Processivity]] is a characteristic of enzymes that function on polymeric substrates. In the case of DNA polymerase, the degree of processivity refers to the average number of nucleotides added each time the enzyme binds a template. The average DNA polymerase requires about one second locating and binding a primer/template junction. Once it is bound, a nonprocessive DNA polymerase adds [[nucleotide]]<nowiki/>s at a rate of one nucleotide per second.<ref name="Losick_2008">{{cite book | author = Losick R, Watson JD, Baker TA, Bell S, Gann A, Levine MW | title = Molecular biology of the gene | publisher = Pearson/Benjamin Cummings | location = San Francisco | year = 2008 | pages = | isbn = 0-8053-9592-X | edition = 6th }}</ref>{{rp|207–208}} Processive DNA polymerases, however, add multiple nucleotides per second, drastically increasing the rate of DNA synthesis. The degree of processivity is directly proportional to the rate of DNA synthesis. The rate of DNA synthesis in a living cell was first determined as the rate of phage T4 DNA elongation in phage infected ''E. coli''. During the period of exponential DNA increase at 37 °C, the rate was 749 nucleotides per second.<ref>{{cite journal |author=McCarthy D, Minner C, Bernstein H, Bernstein C |title=DNA elongation rates and growing point distributions of wild-type phage T4 and a DNA-delay amber mutant |journal=J. Mol. Biol. |volume=106 |issue=4 |pages=963–81 |date=October 1976 |pmid=789903 |doi= 10.1016/0022-2836(76)90346-6|url=}}</ref> |
|||
DNA polymerase’s ability to slide along the DNA template allows increased processivity. There is a dramatic increase in processivity at the [[DNA replication#Replication fork|replication fork]]. This increase is facilitated by the DNA polymerase’s association with proteins known as the sliding [[DNA clamp]]. The clamps are multiple protein subunits associated in the shape of a ring. Using the [[hydrolysis]] of ATP, a class of proteins known as the [[Replication factor C|sliding clamp loading proteins]] open up the ring structure of the sliding DNA clamps allowing binding to and release from the DNA strand. [[Protein-protein interaction]] with the clamp prevents DNA polymerase from diffusing from the DNA template, thereby ensuring that the enzyme binds the same primer/template junction and continues replication.<ref name="Losick_2008"/>{{rp|207–208}} DNA polymerase changes conformation, increasing affinity to the clamp when associated with it and decreasing affinity when it completes the replication of a stretch of DNA to allow release from the clamp. |
|||
==Variation across species== |
|||
{{Infobox protein family |
|||
| Symbol = DNA_pol_A |
|||
| Name = DNA polymerase family A |
|||
| image = PDB 2hht EBI.jpg |
|||
| width = |
|||
| caption = c:o6-methyl-guanine pair in the polymerase-2 basepair position |
|||
| Pfam = PF00476 |
|||
| Pfam_clan = |
|||
| InterPro = IPR001098 |
|||
| SMART = |
|||
-| PROSITE = PDOC00412 |
|||
| MEROPS = |
|||
| SCOP = 1dpi |
|||
| TCDB = |
|||
| OPM family = |
|||
| OPM protein = |
|||
| CAZy = |
|||
| CDD = |
|||
}} |
|||
{{Infobox protein family |
|||
| Symbol = DNA_pol_B |
|||
| Name = DNA polymerase family B |
|||
| image = PDB 2dy4 EBI.jpg |
|||
| width = |
|||
| caption = crystal structure of rb69 gp43 in complex with dna containing thymine glycol |
|||
| Pfam = PF00136 |
|||
| Pfam_clan = CL0194 |
|||
| InterPro = IPR006134 |
|||
| SMART = |
|||
| PROSITE = PDOC00107 |
|||
| MEROPS = |
|||
| SCOP = 1noy |
|||
| TCDB = |
|||
| OPM family = |
|||
| OPM protein = |
|||
| CAZy = |
|||
| CDD = |
|||
}} |
|||
{{Infobox protein family |
|||
| Symbol = DNA_pol_B_2 |
|||
| Name = DNA polymerase type B, organellar and viral |
|||
| image = PDB 1xhz EBI.jpg |
|||
| width = |
|||
| caption = phi29 dna polymerase, orthorhombic crystal form, ssdna complex |
|||
| Pfam = PF03175 |
|||
| Pfam_clan = CL0194 |
|||
| InterPro = IPR004868 |
|||
| SMART = |
|||
| PROSITE = |
|||
| MEROPS = |
|||
| SCOP = |
|||
| TCDB = |
|||
| OPM family = |
|||
| OPM protein = |
|||
| CAZy = |
|||
| CDD = |
|||
}} |
|||
Based on sequence homology, DNA polymerases can be further subdivided into seven different families: A, B, C, D, X, Y, and RT. |
|||
Some [[virus]]<nowiki/>es also encode special DNA polymerases, such as [[Hepatitis B virus DNA polymerase]]. These may selectively replicate viral DNA through a variety of mechanisms. [[Retroviruses]] encode an unusual DNA polymerase called [[reverse transcriptase]], which is an RNA-dependent DNA polymerase (RdDp). It polymerizes DNA from a template of [[RNA]]. |
|||
{| border="1" cellpadding="5" cellspacing="0" style="margin:auto;" |
|||
|- |
|||
! scope="col" style="background:#efefef;" | Family |
|||
! scope="col" style="background:#efefef;" | Types of DNA polymerase |
|||
! scope="col" style="background:#efefef;" | Species |
|||
! scope="col" style="background:#efefef;" | Examples |
|||
|- |
|||
|A |
|||
|Replicative and Repair Polymerases |
|||
|Eukaryotic and Prokaryotic |
|||
|T7 DNA polymerase, Pol I, and DNA Polymerase γ |
|||
|- |
|||
|B |
|||
|Replicative and Repair Polymerases |
|||
|Eukaryotic and Prokaryotic |
|||
|Pol II, Pol B, Pol ζ, Pol α, δ, and ε |
|||
|- |
|||
|C |
|||
|Replicative Polymerases |
|||
|Prokaryotic |
|||
|Pol III |
|||
|- |
|||
|D |
|||
|Replicative Polymerases |
|||
|[[Euryarchaeota]] |
|||
|Not well-characterized |
|||
|- |
|||
|X |
|||
|Replicative and Repair Polymerases |
|||
|Eukaryotic |
|||
|Pol β, Pol σ, Pol λ, Pol μ, and [[Terminal deoxynucleotidyl transferase]] |
|||
|- |
|||
|Y |
|||
|Replicative and Repair Polymerases |
|||
|Eukaryotic and Prokaryotic |
|||
|Pol ι (iota), Pol κ (kappa), Pol IV, and Pol V |
|||
|- |
|||
|RT |
|||
|Replicative and Repair Polymerases |
|||
|Viruses, [[Retroviruses]], and Eukaryotic |
|||
|[[Telomerase]], Hepatitis B virus |
|||
|} |
|||
===Prokaryotic DNA polymerase=== |
|||
====Pol I==== |
|||
Prokaryotic Family A polymerases include the [[DNA polymerase I]] (Pol I) enzyme, which is encoded by the polA gene and ubiquitous among prokaryotes. This repair polymerase is involved in excision repair with 3'-5' and 5'-3' exonuclease activity and processing of [[Okazaki fragment]]s generated during lagging strand synthesis.<ref name="isbn981-4299-16-2">{{cite book | author = Maga G, Hubscher U, Spadari S, Villani G | title = DNA Polymerases: Discovery, Characterization and Functions in Cellular DNA Transactions | publisher = World Scientific Publishing Company | location = | year = 2010 | pages = | isbn = 981-4299-16-2 }}</ref> Pol I is the most abundant polymerase accounting for >95% of polymerase activity in ''E. coli'', yet cells lacking Pol I have been found suggesting Pol I activity can be replaced by the other four polymerases. Pol I adds ~15-20 nucleotides per second, thus showing poor processivity. Instead, Pol I starts adding nucleotides at the RNA primer:template junction known as the origin of replication (ori). Approximately 400 bp downstream from the origin, the Pol III holoenzyme is assembled and takes over replication at a highly processive speed and nature.<ref>{{cite journal |author=Camps M, Loeb LA |title=When pol I goes into high gear: processive DNA synthesis by pol I in the cell |journal=Cell Cycle |volume=3 |issue=2 |pages=116–8 |date=February 2004 |pmid=14712068 |doi= 10.4161/cc.3.2.651 }}</ref> |
|||
====Pol II==== |
|||
[[DNA polymerase II]], a Family B polymerase, is a polB gene product also known as DinA. Pol II has 3'-5' exonuclease activity and participates in DNA repair, replication restart to bypass lesions, and its cell presence can jump from ~30-50 copies per cell to ~200-300 during SOS induction. Pol II is also thought to be a backup to Pol III as it can interact with holoenzyme proteins and assume a high level of processivity. The main role of Pol II is thought to be the ability to direct polymerase activity at the replication fork and helped stalled Pol III bypass terminal mismatches.<ref>{{cite journal |author=Banach-Orlowska M, Fijalkowska IJ, Schaaper RM, Jonczyk P |title=DNA polymerase II as a fidelity factor in chromosomal DNA synthesis in Escherichia coli |journal=Mol. Microbiol. |volume=58 |issue=1 |pages=61–70 |date=October 2005 |pmid=16164549 |doi=10.1111/j.1365-2958.2005.04805.x |url=}}</ref> |
|||
====Pol III==== |
|||
[[DNA polymerase III]] holoenzyme is the primary enzyme involved in DNA replication in ''E. coli'' and belongs to Family C polymerases. It consists of three assemblies: the pol III core, the beta sliding clamp processivity factor and the clamp-loading complex. The core consists of three subunits - α, the polymerase activity hub, ɛ, exonucleolytic proofreader, and θ, which may act as a stabilizer for ɛ. The holoenzyme contains two cores, one for each strand, the lagging and leading.<ref name="pmid16164549">{{cite journal | author = Banach-Orlowska M, Fijalkowska IJ, Schaaper RM, Jonczyk P | title = DNA polymerase II as a fidelity factor in chromosomal DNA synthesis in Escherichia coli | journal = Mol. Microbiol. | volume = 58 | issue = 1 | pages = 61–70 |date=October 2005 | pmid = 16164549 | doi = 10.1111/j.1365-2958.2005.04805.x }}</ref> The beta sliding clamp processivity factor is also present in duplicate, one for each core, to create a clamp that encloses DNA allowing for high processivity.<ref name="pmid7494000">{{cite journal | author = Olson MW, Dallmann HG, McHenry CS | title = DnaX complex of Escherichia coli DNA polymerase III holoenzyme. The chi psi complex functions by increasing the affinity of tau and gamma for delta.delta' to a physiologically relevant range | journal = J. Biol. Chem. | volume = 270 | issue = 49 | pages = 29570–7 |date=December 1995 | pmid = 7494000 | doi = 10.1074/jbc.270.49.29570}}</ref> The third assembly is a seven-subunit (τ2γδδ′χψ) clamp loader complex. Recent research has classified Family C polymerases as a subcategory of Family X with no eukaryotic equivalents.<ref>{{cite web|url=http://www.news-medical.net/health/DNA-Polymerase-Families.aspx |title=DNA Polymerase Families |publisher=News-medical.net |date=2014-05-06 |accessdate=2014-06-28}}</ref> |
|||
====Pol IV==== |
|||
In ''E. coli'', [[DNA polymerase IV]] (Pol 4) is an error-prone DNA polymerase involved in non-targeted mutagenesis.<ref name="pmid12045089">{{cite journal | author = Goodman MF | title = Error-prone repair DNA polymerases in prokaryotes and eukaryotes | journal = Annu. Rev. Biochem. | volume = 71 | issue = | pages = 17–50 | year = 2002 | pmid = 12045089 | doi = 10.1146/annurev.biochem.71.083101.124707 }}</ref> Pol IV is a Family Y polymerase expressed by the dinB gene that is switched on via SOS induction caused by stalled polymerases at the replication fork. During SOS induction, Pol IV production is increased tenfold and one of the functions during this time is to interfere with Pol III holoenzyme processivity. This creates a checkpoint, stops replication, and allows time to repair DNA lesions via the appropriate repair pathway.<ref name="pmid22820381">{{cite journal | author = Mori T | title = Escherichia coli DinB inhibits replication fork progression without significantly inducing the SOS response| journal = Genes Genet Syst. | volume = 87 | issue = 2| pages = 75–87 | year = 2012 | pmid = 22820381 | doi=10.1266/ggs.87.75}}</ref> Another function of Pol IV is to perform [[DNA repair|translesion synthesis]] at the stalled replication fork like, for example, bypassing N2-deoxyguanine adducts at a faster rate than transversing undamaged DNA. Cells lacking dinB gene have a higher rate of mutagenesis caused by DNA damaging agents.<ref name="pmid17377496">{{cite journal | author = Jarosz DF | title = Proficient and accurate bypass of persistent DNA lesions by DinB DNA polymerases.| journal = Cell Cycle | volume = 6 | issue = 7| pages = 817–22 | year = 2007 | pmid = 17377496 | doi=10.4161/cc.6.7.4065}}</ref> |
|||
====Pol V==== |
|||
[[DNA polymerase V]] (Pol V) is a Y-family DNA polymerase that is involved in [[SOS response]] and [[translesion synthesis]] DNA repair mechanisms.<ref name="pmid20441441">{{cite journal | author = Patel M, Jiang Q, Woodgate R, Cox MM, Goodman MF | title = A new model for SOS-induced mutagenesis: how RecA protein activates DNA polymerase V | journal = Crit. Rev. Biochem. Mol. Biol. | volume = 45 | issue = 3 | pages = 171–84 |date=June 2010 | pmid = 20441441 | pmc = 2874081 | doi = 10.3109/10409238.2010.480968 }}</ref> Transcription of Pol V via the umuDC genes is highly regulated to produce only Pol V when damaged DNA is present in the cell generating an SOS response. Stalled polymerases causes RecA to bind to the ssDNA, which causes the LexA protein to autodigest. LexA then loses its ability to repress the transcription of the umuDC operon. The same RecA-ssDNA nucleoprotein posttranslationally modifies the UmuD protein into UmuD' protein. UmuD and UmuD' form a heterodimer that interacts with UmuC, which in turn activates umuC's polymerase catalytic activity on damaged DNA.<ref>{{cite journal |author=Sutton MD, Walker GC |title=Managing DNA polymerases: coordinating DNA replication, DNA repair, and DNA recombination |journal=Proc. Natl. Acad. Sci. U.S.A. |volume=98 |issue=15 |pages=8342–9 |date=July 2001 |pmid=11459973 |pmc=37441 |doi=10.1073/pnas.111036998 |url=}}</ref> |
|||
===Eukaryotic DNA polymerase=== |
|||
====Polymerases β, λ, σ and μ (beta, lambda, sigma, and mu)==== |
|||
Family X polymerases contain the well-known eukaryotic polymerase [[DNA polymerase beta|pol β (beta)]], as well as other eukaryotic polymerases such as Pol σ (sigma), [[DNA polymerase lambda|Pol λ (lambda)]], [[DNA_polymerase_mu|Pol μ (mu)]], and [[terminal deoxynucleotidyl transferase|Terminal deoxynucleotidyl transferase (TdT)]]. Family X polymerases are found mainly in vertebrates, and a few are found in plants and fungi. These polymerases have highly conserved regions that include two helix-hairpin-helix motifs that are imperative in the DNA-polymerase interactions. One motif is located in the 8 kDa domain that interacts with downstream DNA and one motif is located in the thumb domain that interacts with the primer strand. Pol β, encoded by POLB gene, is required for short-patch [[base excision repair]], a DNA repair pathway that is essential for repairing alkylated or oxidized bases as well as [[abasic site]]s. Pol λ and Pol μ, encoded by the [[POLL]] and [[POLM]] genes respectively, are involved in [[non-homologous end-joining]], a mechanism for rejoining DNA double-strand breaks due to hydrogen peroxide and ionizing radiation, respectively. TdT is expressed only in lymphoid tissue, and adds "n nucleotides" to double-strand breaks formed during [[V(D)J recombination]] to promote immunological diversity.<ref name="pmid19631767">{{cite journal | author = Yamtich J, Sweasy JB | title = DNA polymerase family X: function, structure, and cellular roles | journal = Biochim. Biophys. Acta | volume = 1804 | issue = 5 | pages = 1136–50 |date=May 2010 | pmid = 19631767 | pmc = 2846199 | doi = 10.1016/j.bbapap.2009.07.008 }}</ref> |
|||
====Polymerases α, δ and ε (alpha, delta, and epsilon)==== |
|||
[[DNA polymerase alpha|Pol α (alpha)]], [[DNA polymerase delta|Pol δ (delta)]], and [[DNA polymerase epsilon|Pol ε (epsilon)]] are members of Family B Polymerases and are the main polymerases involved with nuclear DNA replication. Pol α complex (pol α-DNA primase complex) consists of four subunits: the catalytic subunit [[POLA1]], the regulatory subunit [[POLA2]], and the small and the large primase subunits [[PRIM1]] and [[PRIM2]] respectively. Once primase has created the RNA primer, Pol α starts replication elongating the primer with ~20 nucleotides.<ref>{{cite book|last=Chansky|first=Michael Lieberman, Allan Marks, Alisa Peet ; illustrations by Matthew|title=Marks' basic medical biochemistry : a clinical approach|year=2012|publisher=Wolter Kluwer Health/Lippincott Williams & Wilkins|location=Philadelphia|isbn=160831572X|page=chapter13|edition=4th}}</ref> Due to its high processivity, Pol δ takes over the leading and lagging strand synthesis from Pol α.<ref name="Losick_2008" />{{rp|218–219}} Pol δ is expressed by genes [[POLD1]], creating the catalytic subunit, [[POLD2]], [[POLD3]], and [[POLD4]] creating the other subunits that interact with [[Proliferating Cell Nuclear Antigen]] (PCNA), which is a [[DNA clamp]] that allows Pol δ to possess processivity.<ref>{{cite journal |author=Chung DW, Zhang JA, Tan CK, Davie EW, So AG, Downey KM |title=Primary structure of the catalytic subunit of human DNA polymerase delta and chromosomal location of the gene |journal=Proc. Natl. Acad. Sci. U.S.A. |volume=88 |issue=24 |pages=11197–201 |date=December 1991 |pmid=1722322 |pmc=53101 |doi= 10.1073/pnas.88.24.11197|url=}}</ref> Pol ε is encoded by the [[POLE1]], the catalytic subunit, [[POLE2]], and [[POLE3]] genes. It was previously thought that Pol ε's main function was to extend the leading strand during replication; however, recent evidence suggests that Pol δ alone replicates the leading and lagging strands of DNA, and Pol ε participates in repairing errors made in the leading strand during Pol δ replication in conjunction with DNA mismatch repair machinery.<ref>{{cite journal |author=Johnson RE, Klassen R, Prakash L, Prakash S |title=A Major Role of DNA Polymerase δ in Replication of Both the Leading and Lagging DNA Strands |journal=Mol. Cell. |volume=59 |issue=2 |pages=163–175 |date=July 16, 2015 |pmid= 26145172 |doi=10.1016/j.molcel.2015.05.038}}</ref> Pol ε's C-terminus region is thought to be essential to cell vitality as well. The C-terminus region is thought to provide a checkpoint before entering anaphase, provide stability to the holoenzyme, and add proteins to the holoenzyme necessary for initiation of replication.<ref>{{cite journal |author=Edwards S, Li CM, Levy DL, Brown J, Snow PM, Campbell JL |title=Saccharomyces cerevisiae DNA polymerase epsilon and polymerase sigma interact physically and functionally, suggesting a role for polymerase epsilon in sister chromatid cohesion |journal=Mol. Cell. Biol. |volume=23 |issue=8 |pages=2733–48 |date=April 2003 |pmid=12665575 |pmc=152548 |doi= 10.1128/mcb.23.8.2733-2748.2003|url=}}</ref> |
|||
====Polymerases η, ι and κ (eta, iota, and kappa)==== |
|||
[[DNA polymerase eta|Pol η (eta)]], Pol ι (iota), and Pol κ (kappa), are Family Y DNA polymerases involved in the DNA repair by translesion synthesis and encoded by genes POLH, [[POLI]], and [[POLK]] respectively. Members of Family Y have five common motifs to aid in binding the substrate and primer terminus and they all include the typical right hand thumb, palm and finger domains with added domains like little finger (LF), polymerase-associated domain (PAD), or wrist. The active site, however, differs between family members due to the different lesions being repaired. Polymerases in Family Y are low-fidelity polymerases, but have been proven to do more good than harm as mutations that affect the polymerase can cause various diseases, such as [[skin cancer]] and [[Xeroderma pigmentosum|Xeroderma Pigmentosum Variant (XPS).]] The importance of these polymerases is evidenced by the fact that gene encoding DNA polymerase η is referred as XPV, because loss of this gene results in the disease Xeroderma Pigmentosum Variant. Pol η is particularly important for allowing accurate translesion synthesis of DNA damage resulting from [[ultraviolet radiation]]. The functionality of Pol κ is not completely understood, but researchers have found two probable functions. Pol κ is thought to act as an extender or an inserter of a specific base at certain DNA lesions. All three translesion synthesis polymerases, along with Rev1, are recruited to damaged lesions via stalled replicative DNA polymerases. There are two pathways of damage repair leading researchers to conclude that the chosen pathway depends on which strand contains the damage, the leading or lagging strand.<ref name="pmid20663485">{{cite journal | author = Ohmori H, Hanafusa T, Ohashi E, Vaziri C | title = Separate roles of structured and unstructured regions of Y-family DNA polymerases | journal = Adv Protein Chem Struct Biol | volume = 78 | issue = | pages = 99–146 | year = 2009 | pmid = 20663485 | pmc = 3103052 | doi = 10.1016/S1876-1623(08)78004-0 }}</ref> |
|||
====Polymerases Rev1 and ζ (zeta)==== |
|||
Pol ζ another B family polymerase, is made of two subunits [[REV3L|Rev3]], the catalytic subunit, and Rev7, which increases the catalytic function of the polymerase, and is involved in translesion synthesis. Pol ζ lacks 3' to 5' exonuclease activity, is unique in that it can extend primers with terminal mismatches. [[REV1|Rev1]] has three regions of interest in the BRCT domain, ubiquitin-binding domain, and C-terminal domain and has dCMP transferase ability, which adds deoxycytidine opposite lesions that would stall replicative polymerases Pol δ and Pol ε. These stalled polymerases activate ubiquitin complexes that in turn disassociate replication polymerases and recruit Pol ζ and Rev1. Together Pol ζ and Rev1 add deoxycytidine and Pol ζ extends past the lesion. Through a yet undetermined process, Pol ζ disassociates and replication polymerases reassociate and continue replication. Pol ζ and Rev1 are not required for replication, but loss of REV3 gene in budding yeast can cause increased sensitivity to DNA-damaging agents due to collapse of replication forks where replication polymerases have stalled.<ref name="pmid18157155">{{cite journal | author = Gan GN, Wittschieben JP, Wittschieben BØ, Wood RD | title = DNA polymerase zeta (pol zeta) in higher eukaryotes | journal = Cell Res. | volume = 18 | issue = 1 | pages = 174–83 |date=January 2008 | pmid = 18157155 | doi = 10.1038/cr.2007.117 }}</ref> |
|||
====Telomerase==== |
|||
[[Telomerase]] is a [[ribonucleoprotein]] recruited to replicate ends of linear chromosomes because normal DNA polymerase cannot replicate the ends, or [[telomere]]. The single-strand 3’ overhang of the double-strand chromosome with the sequence 5’-TTAGGG-3’ recruits telomerase. Telomerase acts like other DNA polymerases by extending the 3’ end, but, unlike other DNA polymerases, telomerase does not require a template. The TERT subunit, an example of a [[reverse transcriptase]], uses the RNA subunit to form the primer–template junction that allows telomerase to extend the 3’ end of chromosome ends. The gradual decrease in size of telomeres as the result of many replications over a lifetime are thought to be associated with the effects of aging.<ref name="Losick_2008"/>{{rp|248–249}} |
|||
====Polymerases γ and θ (gamma and theta)==== |
|||
Pol γ (gamma) and Pol θ (theta) are Family A polymerases. Pol γ, encoded by the [[POLG]] gene, is the only [[Mitochondrial DNA|mtDNA]] polymerase and therefore replicates, repairs, and has proofreading 3'-5' exonuclease and 5' dRP lyase activities. Any mutation that leads to limited or non-functioning Pol γ has a significant effect on mtDNA and is the most common cause of autosomal inherited mitochondrial disorders.<ref name="pmid21732785">{{cite journal | author = Zhang L, Chan SS, Wolff DJ | title = Mitochondrial disorders of DNA polymerase γ dysfunction: from anatomic to molecular pathology diagnosis | journal = Arch. Pathol. Lab. Med. | volume = 135 | issue = 7 | pages = 925–34 |date=July 2011 | pmid = 21732785 | pmc = 3158670 | doi = 10.1043/2010-0356-RAR.1 }}</ref> Pol γ contains a C-terminus polymerase domain and an N-terminus 3'-5' exonuclease domain that are connected via the linker region, which binds the accessory subunit. The accessory subunit binds DNA and is required for processivity of Pol γ. Point mutation A467T in the linker region is responsible for more than one-third of all Pol γ-associated mitochondrial disorders.<ref name="pmid20927567">{{cite journal | author = Stumpf JD, Copeland WC | title = Mitochondrial DNA replication and disease: insights from DNA polymerase γ mutations | journal = Cell. Mol. Life Sci. | volume = 68 | issue = 2 | pages = 219–33 |date=January 2011 | pmid = 20927567 | pmc = 3046768 | doi = 10.1007/s00018-010-0530-4 }}</ref> While many homologs of Pol θ, encoded by the [[POLQ]] gene, are found in eukaryotes, its function is not clearly understood. The sequence of amino acids in the C-terminus is what classifies Pol θ as Family A polymerase, although the error rate for Pol θ is more closely related to Family Y polymerases. Pol θ extends mismatched primer termini and can bypass abasic sites by adding a nucleotide. It also has Deoxyribophosphodiesterase (dRPase) activity in the polymerase domain and can show [[ATPase]] activity in close proximity to ssDNA.<ref name="pmid22135286">{{cite journal | author = Hogg M, Sauer-Eriksson AE, Johansson E | title = Promiscuous DNA synthesis by human DNA polymerase θ | journal = Nucleic Acids Res. | volume = 40 | issue = 6 | pages = 2611–22 |date=March 2012 | pmid = 22135286 | pmc = 3315306 | doi = 10.1093/nar/gkr1102 }}</ref> |
|||
====Polymerase ν (nu)==== |
|||
{{see|DNA polymerase nu}} |
|||
====Reverse transcriptase==== |
|||
Retroviruses encode an unusual DNA polymerase called [[reverse transcriptase]], which is an RNA-dependent DNA polymerase (RdDp) that synthesizes DNA from a template of RNA. The reverse transcriptase family contain both DNA polymerase functionality and RNase H functionality, which degrades RNA base-paired to DNA. Some retrovirus examples include [[Hepatitis B]] virus and [[HIV]].<ref name="Losick_2008"/>{{rp|}} |
|||
==See also== |
|||
*[[Molecular machine#Biological|Biological machines]] |
|||
*[[DNA repair]] |
|||
*[[DNA replication]] |
|||
*[[DNA sequencing]] |
|||
*[[Enzyme catalysis]] |
|||
*[[Genetic recombination]] |
|||
*[[Molecular cloning]] |
|||
*[[Polymerase chain reaction]] |
|||
*[[Protein dynamics#Global flexibility: multiple domains|Protein domain dynamics]] |
|||
*[[Reverse transcription]] |
|||
*[[RNA polymerase]] |
|||
*[[Taq DNA polymerase]] |
|||
== References == |
|||
{{reflist|2}} |
|||
== Further reading == |
|||
{{refbegin}} |
|||
* {{cite journal | author = Burgers PM, Koonin EV, Bruford E, Blanco L, Burtis KC, Christman MF, Copeland WC, Friedberg EC, Hanaoka F, Hinkle DC, Lawrence CW, Nakanishi M, Ohmori H, Prakash L, Prakash S, Reynaud CA, Sugino A, Todo T, Wang Z, Weill JC, Woodgate R | title = Eukaryotic DNA polymerases: proposal for a revised nomenclature | journal = J. Biol. Chem. | volume = 276 | issue = 47 | pages = 43487–90 |date=November 2001 | pmid = 11579108 | doi = 10.1074/jbc.R100056200 }} |
|||
{{refend}} |
|||
==External links== |
|||
{{Commons category|DNA polymerases}} |
|||
* {{MeshName|DNA+polymerases}} |
|||
* {{PDB Molecule of the Month|3|DNA polymerase}} |
|||
* [http://researchnews.osu.edu/archive/repprot.htm Unusual repair mechanism in DNA polymerase lambda], [[Ohio State University]], July 25, 2006. |
|||
* [http://wehi.edu.au/education/wehitv/molecular_visualisations_of_dna/ A great animation of DNA Polymerase from WEHI at 1:45 minutes in] |
|||
* [http://www.pdbe.org/emsearch/dna\%20polymerase 3D macromolecular structures of DNA polymerase from the EM Data Bank(EMDB)] |
|||
{{DNA replication}} |
|||
{{Kinases}} |
|||
{{Enzymes}} |
|||
{{Portal bar|Molecular and Cellular Biology|border=no}} |
|||
[[Category:EC 2.7.7]] |
|||
[[Category:DNA replication]] |
|||
[[Category:DNA]] |
|||
[[Category:Enzymes]] |
|||
Phiên bản lúc 07:31, ngày 4 tháng 4 năm 2016
| DNA-directed DNA polymerase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 3D structure of the DNA-binding helix-turn-helix motifs in human DNA polymerase beta (based on PDB file 7ICG) | |||||||||
| Mã định danh (ID) | |||||||||
| Mã EC | 2.7.7.7 | ||||||||
| Mã CAS | 9012-90-2 | ||||||||
| Các dữ liệu thông tin | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | chu trình chuyển hóa | ||||||||
| PRIAM | profile | ||||||||
| Các cấu trúc PDB | RCSB PDB PDBj PDBe PDBsum | ||||||||
| Bản thể gen | AmiGO / EGO | ||||||||
| |||||||||
The DNA polymerases are enzymes that create DNA molecules by assembling nucleotides, the building blocks of DNA. These enzymes are essential to DNA replication and usually work in pairs to create two identical DNA strands from a single original DNA molecule. During this process, DNA polymerase “reads” the existing DNA strands to create two new strands that match the existing ones. [1][2][3][4][5][6]
This enzyme catalyses the following chemical reaction
- deoxynucleoside triphosphate + DNAn diphosphate + DNAn+1
Catalyses DNA-template-directed extension of the 3'- end of a DNA strand by one nucleotide at a time.
Every time a cell divides, DNA polymerase is required to help duplicate the cell’s DNA, so that a copy of the original DNA molecule can be passed to each of the daughter cells. In this way, genetic information is transmitted from generation to generation.
Before replication can take place, an enzyme called helicase unwinds the DNA molecule from its tightly woven form. This opens up or “unzips” the double-stranded DNA to give two single strands of DNA that can be used as templates for replication.
History
In 1956, Arthur Kornberg and colleagues discovered the enzyme DNA polymerase I, also known as Pol I, in Escherichia coli. They described the DNA replication process by which DNA polymerase copies the base sequence of a template DNA strand. Subsequently, in 1959, Kornberg was awarded the Nobel Prize in Physiology or Medicine for this work.[7] DNA polymerase II was also discovered by Kornberg and Malcolm E. Gefter in 1970 while further elucidating the role of Pol I in E. coli DNA replication.[8]
Function

The main function of DNA polymerase is to make DNA from nucleotides, the building blocks of DNA. The DNA copies are created by the pairing of nucleotides to bases present on each strand of the original DNA molecule. This pairing always occurs in specific combinations, with cytosine along with guanine, and thymine along with adenine, forming two separate pairs, respectively.
When creating DNA, DNA polymerase can add free nucleotides only to the 3' end of the newly forming strand. This results in elongation of the newly forming strand in a 5'-3' direction. No known DNA polymerase is able to begin a new chain (de novo); it can only add a nucleotide onto a pre-existing 3'-OH group, and therefore needs a primer at which it can add the first nucleotide. Primers consist of RNA or DNA bases (or both). In DNA replication, the first two bases are always RNA, and are synthesized by another enzyme called primase. Enzymes,helicase and topoisomerase II, are required to unwind DNA from a double-strand structure to a single-strand structure to facilitate replication of each strand consistent with the semiconservative model of DNA replication.
It is important to note that the directionality of the newly forming strand (the daughter strand) is opposite to the direction in which DNA polymerase moves along the template strand. Since DNA polymerase requires a free 3' OH group for initiation of synthesis, it can synthesize in only one direction by extending the 3' end of the preexisting nucleotide chain. Hence, DNA polymerase moves along the template strand in a 3'-5' direction, and the daughter strand is formed in a 5'-3' direction. This difference enables the resultant double-strand DNA formed to be composed of two DNA strands that are antiparallel to each other.
The function of DNA polymerase is not quite perfect, with the enzyme making about one mistake for every billion base pairs copied. Error correction is a property of some, but not all, DNA polymerases. This process corrects mistakes in newly synthesized DNA. When an incorrect base pair is recognized, DNA polymerase moves backwards by one base pair of DNA. The 3'-5' exonuclease activity of the enzyme allows the incorrect base pair to be excised (this activity is known as proofreading). Following base excision, the polymerase can re-insert the correct base and replication can continue forwards. This preserves the integrity of the original DNA strand that is passed onto the daughter cells.
Structure
The known DNA polymerases have highly conserved structure, which means that their overall catalytic subunits vary very little from species to species, independent of their domain structures. Conserved structures usually indicate important, irreplaceable functions of the cell, the maintenance of which provides evolutionary advantages. The shape can be described as resembling a right hand with thumb, finger, and palm domains. The palm domain appears to function in catalyzing the transfer of phosphoryl groups in the phosphoryl transfer reaction. DNA is bound to the palm when the enzyme is active. This reaction is believed to be catalyzed by a two-metal-ion mechanism. The finger domain functions to bind the nucleotide triphosphate with the template base. The thumb domain plays a potential role in the processivity, translocation, and positioning of the DNA.[9]
Processivity
DNA polymerase’s rapid catalysis is due to its processive nature. Processivity is a characteristic of enzymes that function on polymeric substrates. In the case of DNA polymerase, the degree of processivity refers to the average number of nucleotides added each time the enzyme binds a template. The average DNA polymerase requires about one second locating and binding a primer/template junction. Once it is bound, a nonprocessive DNA polymerase adds nucleotides at a rate of one nucleotide per second.[10]:207–208 Processive DNA polymerases, however, add multiple nucleotides per second, drastically increasing the rate of DNA synthesis. The degree of processivity is directly proportional to the rate of DNA synthesis. The rate of DNA synthesis in a living cell was first determined as the rate of phage T4 DNA elongation in phage infected E. coli. During the period of exponential DNA increase at 37 °C, the rate was 749 nucleotides per second.[11]
DNA polymerase’s ability to slide along the DNA template allows increased processivity. There is a dramatic increase in processivity at the replication fork. This increase is facilitated by the DNA polymerase’s association with proteins known as the sliding DNA clamp. The clamps are multiple protein subunits associated in the shape of a ring. Using the hydrolysis of ATP, a class of proteins known as the sliding clamp loading proteins open up the ring structure of the sliding DNA clamps allowing binding to and release from the DNA strand. Protein-protein interaction with the clamp prevents DNA polymerase from diffusing from the DNA template, thereby ensuring that the enzyme binds the same primer/template junction and continues replication.[10]:207–208 DNA polymerase changes conformation, increasing affinity to the clamp when associated with it and decreasing affinity when it completes the replication of a stretch of DNA to allow release from the clamp.
Variation across species
| DNA polymerase family A | |||||||||
|---|---|---|---|---|---|---|---|---|---|

| |||||||||
| c:o6-methyl-guanine pair in the polymerase-2 basepair position | |||||||||
| Danh pháp | |||||||||
| Ký hiệu | DNA_pol_A | ||||||||
| Pfam | PF00476 | ||||||||
| InterPro | IPR001098 | ||||||||
| SMART | - | ||||||||
| PROSITE | PDOC00412 | ||||||||
| SCOP | 1dpi | ||||||||
| |||||||||
| DNA polymerase family B | |||||||||
|---|---|---|---|---|---|---|---|---|---|

| |||||||||
| crystal structure of rb69 gp43 in complex with dna containing thymine glycol | |||||||||
| Danh pháp | |||||||||
| Ký hiệu | DNA_pol_B | ||||||||
| Pfam | PF00136 | ||||||||
| Pfam clan | CL0194 | ||||||||
| InterPro | IPR006134 | ||||||||
| PROSITE | PDOC00107 | ||||||||
| SCOP | 1noy | ||||||||
| |||||||||
| DNA polymerase type B, organellar and viral | |||||||||
|---|---|---|---|---|---|---|---|---|---|

| |||||||||
| phi29 dna polymerase, orthorhombic crystal form, ssdna complex | |||||||||
| Danh pháp | |||||||||
| Ký hiệu | DNA_pol_B_2 | ||||||||
| Pfam | PF03175 | ||||||||
| Pfam clan | CL0194 | ||||||||
| InterPro | IPR004868 | ||||||||
| |||||||||
Based on sequence homology, DNA polymerases can be further subdivided into seven different families: A, B, C, D, X, Y, and RT.
Some viruses also encode special DNA polymerases, such as Hepatitis B virus DNA polymerase. These may selectively replicate viral DNA through a variety of mechanisms. Retroviruses encode an unusual DNA polymerase called reverse transcriptase, which is an RNA-dependent DNA polymerase (RdDp). It polymerizes DNA from a template of RNA.
| Family | Types of DNA polymerase | Species | Examples |
|---|---|---|---|
| A | Replicative and Repair Polymerases | Eukaryotic and Prokaryotic | T7 DNA polymerase, Pol I, and DNA Polymerase γ |
| B | Replicative and Repair Polymerases | Eukaryotic and Prokaryotic | Pol II, Pol B, Pol ζ, Pol α, δ, and ε |
| C | Replicative Polymerases | Prokaryotic | Pol III |
| D | Replicative Polymerases | Euryarchaeota | Not well-characterized |
| X | Replicative and Repair Polymerases | Eukaryotic | Pol β, Pol σ, Pol λ, Pol μ, and Terminal deoxynucleotidyl transferase |
| Y | Replicative and Repair Polymerases | Eukaryotic and Prokaryotic | Pol ι (iota), Pol κ (kappa), Pol IV, and Pol V |
| RT | Replicative and Repair Polymerases | Viruses, Retroviruses, and Eukaryotic | Telomerase, Hepatitis B virus |
Prokaryotic DNA polymerase
Pol I
Prokaryotic Family A polymerases include the DNA polymerase I (Pol I) enzyme, which is encoded by the polA gene and ubiquitous among prokaryotes. This repair polymerase is involved in excision repair with 3'-5' and 5'-3' exonuclease activity and processing of Okazaki fragments generated during lagging strand synthesis.[12] Pol I is the most abundant polymerase accounting for >95% of polymerase activity in E. coli, yet cells lacking Pol I have been found suggesting Pol I activity can be replaced by the other four polymerases. Pol I adds ~15-20 nucleotides per second, thus showing poor processivity. Instead, Pol I starts adding nucleotides at the RNA primer:template junction known as the origin of replication (ori). Approximately 400 bp downstream from the origin, the Pol III holoenzyme is assembled and takes over replication at a highly processive speed and nature.[13]
Pol II
DNA polymerase II, a Family B polymerase, is a polB gene product also known as DinA. Pol II has 3'-5' exonuclease activity and participates in DNA repair, replication restart to bypass lesions, and its cell presence can jump from ~30-50 copies per cell to ~200-300 during SOS induction. Pol II is also thought to be a backup to Pol III as it can interact with holoenzyme proteins and assume a high level of processivity. The main role of Pol II is thought to be the ability to direct polymerase activity at the replication fork and helped stalled Pol III bypass terminal mismatches.[14]
Pol III
DNA polymerase III holoenzyme is the primary enzyme involved in DNA replication in E. coli and belongs to Family C polymerases. It consists of three assemblies: the pol III core, the beta sliding clamp processivity factor and the clamp-loading complex. The core consists of three subunits - α, the polymerase activity hub, ɛ, exonucleolytic proofreader, and θ, which may act as a stabilizer for ɛ. The holoenzyme contains two cores, one for each strand, the lagging and leading.[15] The beta sliding clamp processivity factor is also present in duplicate, one for each core, to create a clamp that encloses DNA allowing for high processivity.[16] The third assembly is a seven-subunit (τ2γδδ′χψ) clamp loader complex. Recent research has classified Family C polymerases as a subcategory of Family X with no eukaryotic equivalents.[17]
Pol IV
In E. coli, DNA polymerase IV (Pol 4) is an error-prone DNA polymerase involved in non-targeted mutagenesis.[18] Pol IV is a Family Y polymerase expressed by the dinB gene that is switched on via SOS induction caused by stalled polymerases at the replication fork. During SOS induction, Pol IV production is increased tenfold and one of the functions during this time is to interfere with Pol III holoenzyme processivity. This creates a checkpoint, stops replication, and allows time to repair DNA lesions via the appropriate repair pathway.[19] Another function of Pol IV is to perform translesion synthesis at the stalled replication fork like, for example, bypassing N2-deoxyguanine adducts at a faster rate than transversing undamaged DNA. Cells lacking dinB gene have a higher rate of mutagenesis caused by DNA damaging agents.[20]
Pol V
DNA polymerase V (Pol V) is a Y-family DNA polymerase that is involved in SOS response and translesion synthesis DNA repair mechanisms.[21] Transcription of Pol V via the umuDC genes is highly regulated to produce only Pol V when damaged DNA is present in the cell generating an SOS response. Stalled polymerases causes RecA to bind to the ssDNA, which causes the LexA protein to autodigest. LexA then loses its ability to repress the transcription of the umuDC operon. The same RecA-ssDNA nucleoprotein posttranslationally modifies the UmuD protein into UmuD' protein. UmuD and UmuD' form a heterodimer that interacts with UmuC, which in turn activates umuC's polymerase catalytic activity on damaged DNA.[22]
Eukaryotic DNA polymerase
Polymerases β, λ, σ and μ (beta, lambda, sigma, and mu)
Family X polymerases contain the well-known eukaryotic polymerase pol β (beta), as well as other eukaryotic polymerases such as Pol σ (sigma), Pol λ (lambda), Pol μ (mu), and Terminal deoxynucleotidyl transferase (TdT). Family X polymerases are found mainly in vertebrates, and a few are found in plants and fungi. These polymerases have highly conserved regions that include two helix-hairpin-helix motifs that are imperative in the DNA-polymerase interactions. One motif is located in the 8 kDa domain that interacts with downstream DNA and one motif is located in the thumb domain that interacts with the primer strand. Pol β, encoded by POLB gene, is required for short-patch base excision repair, a DNA repair pathway that is essential for repairing alkylated or oxidized bases as well as abasic sites. Pol λ and Pol μ, encoded by the POLL and POLM genes respectively, are involved in non-homologous end-joining, a mechanism for rejoining DNA double-strand breaks due to hydrogen peroxide and ionizing radiation, respectively. TdT is expressed only in lymphoid tissue, and adds "n nucleotides" to double-strand breaks formed during V(D)J recombination to promote immunological diversity.[23]
Polymerases α, δ and ε (alpha, delta, and epsilon)
Pol α (alpha), Pol δ (delta), and Pol ε (epsilon) are members of Family B Polymerases and are the main polymerases involved with nuclear DNA replication. Pol α complex (pol α-DNA primase complex) consists of four subunits: the catalytic subunit POLA1, the regulatory subunit POLA2, and the small and the large primase subunits PRIM1 and PRIM2 respectively. Once primase has created the RNA primer, Pol α starts replication elongating the primer with ~20 nucleotides.[24] Due to its high processivity, Pol δ takes over the leading and lagging strand synthesis from Pol α.[10]:218–219 Pol δ is expressed by genes POLD1, creating the catalytic subunit, POLD2, POLD3, and POLD4 creating the other subunits that interact with Proliferating Cell Nuclear Antigen (PCNA), which is a DNA clamp that allows Pol δ to possess processivity.[25] Pol ε is encoded by the POLE1, the catalytic subunit, POLE2, and POLE3 genes. It was previously thought that Pol ε's main function was to extend the leading strand during replication; however, recent evidence suggests that Pol δ alone replicates the leading and lagging strands of DNA, and Pol ε participates in repairing errors made in the leading strand during Pol δ replication in conjunction with DNA mismatch repair machinery.[26] Pol ε's C-terminus region is thought to be essential to cell vitality as well. The C-terminus region is thought to provide a checkpoint before entering anaphase, provide stability to the holoenzyme, and add proteins to the holoenzyme necessary for initiation of replication.[27]
Polymerases η, ι and κ (eta, iota, and kappa)
Pol η (eta), Pol ι (iota), and Pol κ (kappa), are Family Y DNA polymerases involved in the DNA repair by translesion synthesis and encoded by genes POLH, POLI, and POLK respectively. Members of Family Y have five common motifs to aid in binding the substrate and primer terminus and they all include the typical right hand thumb, palm and finger domains with added domains like little finger (LF), polymerase-associated domain (PAD), or wrist. The active site, however, differs between family members due to the different lesions being repaired. Polymerases in Family Y are low-fidelity polymerases, but have been proven to do more good than harm as mutations that affect the polymerase can cause various diseases, such as skin cancer and Xeroderma Pigmentosum Variant (XPS). The importance of these polymerases is evidenced by the fact that gene encoding DNA polymerase η is referred as XPV, because loss of this gene results in the disease Xeroderma Pigmentosum Variant. Pol η is particularly important for allowing accurate translesion synthesis of DNA damage resulting from ultraviolet radiation. The functionality of Pol κ is not completely understood, but researchers have found two probable functions. Pol κ is thought to act as an extender or an inserter of a specific base at certain DNA lesions. All three translesion synthesis polymerases, along with Rev1, are recruited to damaged lesions via stalled replicative DNA polymerases. There are two pathways of damage repair leading researchers to conclude that the chosen pathway depends on which strand contains the damage, the leading or lagging strand.[28]
Polymerases Rev1 and ζ (zeta)
Pol ζ another B family polymerase, is made of two subunits Rev3, the catalytic subunit, and Rev7, which increases the catalytic function of the polymerase, and is involved in translesion synthesis. Pol ζ lacks 3' to 5' exonuclease activity, is unique in that it can extend primers with terminal mismatches. Rev1 has three regions of interest in the BRCT domain, ubiquitin-binding domain, and C-terminal domain and has dCMP transferase ability, which adds deoxycytidine opposite lesions that would stall replicative polymerases Pol δ and Pol ε. These stalled polymerases activate ubiquitin complexes that in turn disassociate replication polymerases and recruit Pol ζ and Rev1. Together Pol ζ and Rev1 add deoxycytidine and Pol ζ extends past the lesion. Through a yet undetermined process, Pol ζ disassociates and replication polymerases reassociate and continue replication. Pol ζ and Rev1 are not required for replication, but loss of REV3 gene in budding yeast can cause increased sensitivity to DNA-damaging agents due to collapse of replication forks where replication polymerases have stalled.[29]
Telomerase
Telomerase is a ribonucleoprotein recruited to replicate ends of linear chromosomes because normal DNA polymerase cannot replicate the ends, or telomere. The single-strand 3’ overhang of the double-strand chromosome with the sequence 5’-TTAGGG-3’ recruits telomerase. Telomerase acts like other DNA polymerases by extending the 3’ end, but, unlike other DNA polymerases, telomerase does not require a template. The TERT subunit, an example of a reverse transcriptase, uses the RNA subunit to form the primer–template junction that allows telomerase to extend the 3’ end of chromosome ends. The gradual decrease in size of telomeres as the result of many replications over a lifetime are thought to be associated with the effects of aging.[10]:248–249
Polymerases γ and θ (gamma and theta)
Pol γ (gamma) and Pol θ (theta) are Family A polymerases. Pol γ, encoded by the POLG gene, is the only mtDNA polymerase and therefore replicates, repairs, and has proofreading 3'-5' exonuclease and 5' dRP lyase activities. Any mutation that leads to limited or non-functioning Pol γ has a significant effect on mtDNA and is the most common cause of autosomal inherited mitochondrial disorders.[30] Pol γ contains a C-terminus polymerase domain and an N-terminus 3'-5' exonuclease domain that are connected via the linker region, which binds the accessory subunit. The accessory subunit binds DNA and is required for processivity of Pol γ. Point mutation A467T in the linker region is responsible for more than one-third of all Pol γ-associated mitochondrial disorders.[31] While many homologs of Pol θ, encoded by the POLQ gene, are found in eukaryotes, its function is not clearly understood. The sequence of amino acids in the C-terminus is what classifies Pol θ as Family A polymerase, although the error rate for Pol θ is more closely related to Family Y polymerases. Pol θ extends mismatched primer termini and can bypass abasic sites by adding a nucleotide. It also has Deoxyribophosphodiesterase (dRPase) activity in the polymerase domain and can show ATPase activity in close proximity to ssDNA.[32]
Polymerase ν (nu)
Reverse transcriptase
Retroviruses encode an unusual DNA polymerase called reverse transcriptase, which is an RNA-dependent DNA polymerase (RdDp) that synthesizes DNA from a template of RNA. The reverse transcriptase family contain both DNA polymerase functionality and RNase H functionality, which degrades RNA base-paired to DNA. Some retrovirus examples include Hepatitis B virus and HIV.[10]:
See also
- Biological machines
- DNA repair
- DNA replication
- DNA sequencing
- Enzyme catalysis
- Genetic recombination
- Molecular cloning
- Polymerase chain reaction
- Protein domain dynamics
- Reverse transcription
- RNA polymerase
- Taq DNA polymerase
References
- ^ Bollum, F.J. (1960). “Calf thymus polymerase”. J. Biol. Chem. 235: 2399–2403. PMID 13802334.
- ^ Falaschi, A. and Kornberg, A. (1966). “Biochemical studies of bacterial sporulation. II. Deoxy-ribonucleic acid polymerase in spores of Bacillus subtilis”. J. Biol. Chem. 241: 1478–1482. PMID 4957767.Quản lý CS1: nhiều tên: danh sách tác giả (liên kết)
- ^ Lehman, I.R., Bessman, M.J., Simms, E.S. and Kornberg, A. (1958). “Enzymatic synthesis of deoxyribonucleic acid. I. Preparation of substrates and partial purification of an enzyme from Escherichia coli”. J. Biol. Chem. 233: 163–170. PMID 13563462.Quản lý CS1: nhiều tên: danh sách tác giả (liên kết)
- ^ Richardson, C.C., Schildkraut, C.L., Aposhian, H.V. and Kornberg, A. (1964). “Enzymatic synthesis of deoxyribonucleic acid. XIV. Further purification and properties of deoxyribonucleic acid polymerase of Escherichia coli”. J. Biol. Chem. 239: 222–232. PMID 14114848.Quản lý CS1: nhiều tên: danh sách tác giả (liên kết)
- ^ Schachman, H.K., Adler, J., Radding, C.M., Lehman, I.R. and Kornberg, A. (1960). “Enzymatic synthesis of deoxyribonucleic acid. VII. Synthesis of a polymer of deoxyadenylate and deoxythymidylate”. J. Biol. Chem. 235: 3242–3249. PMID 13747134.Quản lý CS1: nhiều tên: danh sách tác giả (liên kết)
- ^ Zimmerman, B.K. (1966). “Purification and properties of deoxyribonucleic acid polymerase from Micrococcus lysodeikticus”. J. Biol. Chem. 241: 2035–2041. PMID 5946628.
- ^ “The Nobel Prize in Physiology or Medicine 1959”. Nobel Foundation. Truy cập ngày 1 tháng 12 năm 2012.
- ^ Tessman I, Kennedy MA (tháng 2 năm 1994). “DNA polymerase II of Escherichia coli in the bypass of abasic sites in vivo”. Genetics. 136 (2): 439–48. PMC 1205799. PMID 7908652.
- ^ Steitz TA (tháng 6 năm 1999). “DNA polymerases: structural diversity and common mechanisms”. J. Biol. Chem. 274 (25): 17395–8. doi:10.1074/jbc.274.25.17395. PMID 10364165.
- ^ a b c d e Losick R, Watson JD, Baker TA, Bell S, Gann A, Levine MW (2008). Molecular biology of the gene (ấn bản 6). San Francisco: Pearson/Benjamin Cummings. ISBN 0-8053-9592-X.Quản lý CS1: nhiều tên: danh sách tác giả (liên kết)
- ^ McCarthy D, Minner C, Bernstein H, Bernstein C (tháng 10 năm 1976). “DNA elongation rates and growing point distributions of wild-type phage T4 and a DNA-delay amber mutant”. J. Mol. Biol. 106 (4): 963–81. doi:10.1016/0022-2836(76)90346-6. PMID 789903.Quản lý CS1: nhiều tên: danh sách tác giả (liên kết)
- ^ Maga G, Hubscher U, Spadari S, Villani G (2010). DNA Polymerases: Discovery, Characterization and Functions in Cellular DNA Transactions. World Scientific Publishing Company. ISBN 981-4299-16-2.Quản lý CS1: nhiều tên: danh sách tác giả (liên kết)
- ^ Camps M, Loeb LA (tháng 2 năm 2004). “When pol I goes into high gear: processive DNA synthesis by pol I in the cell”. Cell Cycle. 3 (2): 116–8. doi:10.4161/cc.3.2.651. PMID 14712068.
- ^ Banach-Orlowska M, Fijalkowska IJ, Schaaper RM, Jonczyk P (tháng 10 năm 2005). “DNA polymerase II as a fidelity factor in chromosomal DNA synthesis in Escherichia coli”. Mol. Microbiol. 58 (1): 61–70. doi:10.1111/j.1365-2958.2005.04805.x. PMID 16164549.Quản lý CS1: nhiều tên: danh sách tác giả (liên kết)
- ^ Banach-Orlowska M, Fijalkowska IJ, Schaaper RM, Jonczyk P (tháng 10 năm 2005). “DNA polymerase II as a fidelity factor in chromosomal DNA synthesis in Escherichia coli”. Mol. Microbiol. 58 (1): 61–70. doi:10.1111/j.1365-2958.2005.04805.x. PMID 16164549.Quản lý CS1: nhiều tên: danh sách tác giả (liên kết)
- ^ Olson MW, Dallmann HG, McHenry CS (tháng 12 năm 1995). “DnaX complex of Escherichia coli DNA polymerase III holoenzyme. The chi psi complex functions by increasing the affinity of tau and gamma for delta.delta' to a physiologically relevant range”. J. Biol. Chem. 270 (49): 29570–7. doi:10.1074/jbc.270.49.29570. PMID 7494000.Quản lý CS1: nhiều tên: danh sách tác giả (liên kết)
- ^ “DNA Polymerase Families”. News-medical.net. 6 tháng 5 năm 2014. Truy cập ngày 28 tháng 6 năm 2014.
- ^ Goodman MF (2002). “Error-prone repair DNA polymerases in prokaryotes and eukaryotes”. Annu. Rev. Biochem. 71: 17–50. doi:10.1146/annurev.biochem.71.083101.124707. PMID 12045089.
- ^ Mori T (2012). “Escherichia coli DinB inhibits replication fork progression without significantly inducing the SOS response”. Genes Genet Syst. 87 (2): 75–87. doi:10.1266/ggs.87.75. PMID 22820381.
- ^ Jarosz DF (2007). “Proficient and accurate bypass of persistent DNA lesions by DinB DNA polymerases”. Cell Cycle. 6 (7): 817–22. doi:10.4161/cc.6.7.4065. PMID 17377496.
- ^ Patel M, Jiang Q, Woodgate R, Cox MM, Goodman MF (tháng 6 năm 2010). “A new model for SOS-induced mutagenesis: how RecA protein activates DNA polymerase V”. Crit. Rev. Biochem. Mol. Biol. 45 (3): 171–84. doi:10.3109/10409238.2010.480968. PMC 2874081. PMID 20441441.Quản lý CS1: nhiều tên: danh sách tác giả (liên kết)
- ^ Sutton MD, Walker GC (tháng 7 năm 2001). “Managing DNA polymerases: coordinating DNA replication, DNA repair, and DNA recombination”. Proc. Natl. Acad. Sci. U.S.A. 98 (15): 8342–9. doi:10.1073/pnas.111036998. PMC 37441. PMID 11459973.
- ^ Yamtich J, Sweasy JB (tháng 5 năm 2010). “DNA polymerase family X: function, structure, and cellular roles”. Biochim. Biophys. Acta. 1804 (5): 1136–50. doi:10.1016/j.bbapap.2009.07.008. PMC 2846199. PMID 19631767.
- ^ Chansky, Michael Lieberman, Allan Marks, Alisa Peet ; illustrations by Matthew (2012). Marks' basic medical biochemistry : a clinical approach (ấn bản 4). Philadelphia: Wolter Kluwer Health/Lippincott Williams & Wilkins. tr. chapter13. ISBN 160831572X.
- ^ Chung DW, Zhang JA, Tan CK, Davie EW, So AG, Downey KM (tháng 12 năm 1991). “Primary structure of the catalytic subunit of human DNA polymerase delta and chromosomal location of the gene”. Proc. Natl. Acad. Sci. U.S.A. 88 (24): 11197–201. doi:10.1073/pnas.88.24.11197. PMC 53101. PMID 1722322.Quản lý CS1: nhiều tên: danh sách tác giả (liên kết)
- ^ Johnson RE, Klassen R, Prakash L, Prakash S (16 tháng 7 năm 2015). “A Major Role of DNA Polymerase δ in Replication of Both the Leading and Lagging DNA Strands”. Mol. Cell. 59 (2): 163–175. doi:10.1016/j.molcel.2015.05.038. PMID 26145172.Quản lý CS1: nhiều tên: danh sách tác giả (liên kết)
- ^ Edwards S, Li CM, Levy DL, Brown J, Snow PM, Campbell JL (tháng 4 năm 2003). “Saccharomyces cerevisiae DNA polymerase epsilon and polymerase sigma interact physically and functionally, suggesting a role for polymerase epsilon in sister chromatid cohesion”. Mol. Cell. Biol. 23 (8): 2733–48. doi:10.1128/mcb.23.8.2733-2748.2003. PMC 152548. PMID 12665575.Quản lý CS1: nhiều tên: danh sách tác giả (liên kết)
- ^ Ohmori H, Hanafusa T, Ohashi E, Vaziri C (2009). “Separate roles of structured and unstructured regions of Y-family DNA polymerases”. Adv Protein Chem Struct Biol. 78: 99–146. doi:10.1016/S1876-1623(08)78004-0. PMC 3103052. PMID 20663485.Quản lý CS1: nhiều tên: danh sách tác giả (liên kết)
- ^ Gan GN, Wittschieben JP, Wittschieben BØ, Wood RD (tháng 1 năm 2008). “DNA polymerase zeta (pol zeta) in higher eukaryotes”. Cell Res. 18 (1): 174–83. doi:10.1038/cr.2007.117. PMID 18157155.Quản lý CS1: nhiều tên: danh sách tác giả (liên kết)
- ^ Zhang L, Chan SS, Wolff DJ (tháng 7 năm 2011). “Mitochondrial disorders of DNA polymerase γ dysfunction: from anatomic to molecular pathology diagnosis”. Arch. Pathol. Lab. Med. 135 (7): 925–34. doi:10.1043/2010-0356-RAR.1. PMC 3158670. PMID 21732785.Quản lý CS1: nhiều tên: danh sách tác giả (liên kết)
- ^ Stumpf JD, Copeland WC (tháng 1 năm 2011). “Mitochondrial DNA replication and disease: insights from DNA polymerase γ mutations”. Cell. Mol. Life Sci. 68 (2): 219–33. doi:10.1007/s00018-010-0530-4. PMC 3046768. PMID 20927567.
- ^ Hogg M, Sauer-Eriksson AE, Johansson E (tháng 3 năm 2012). “Promiscuous DNA synthesis by human DNA polymerase θ”. Nucleic Acids Res. 40 (6): 2611–22. doi:10.1093/nar/gkr1102. PMC 3315306. PMID 22135286.Quản lý CS1: nhiều tên: danh sách tác giả (liên kết)
Further reading
- Burgers PM, Koonin EV, Bruford E, Blanco L, Burtis KC, Christman MF, Copeland WC, Friedberg EC, Hanaoka F, Hinkle DC, Lawrence CW, Nakanishi M, Ohmori H, Prakash L, Prakash S, Reynaud CA, Sugino A, Todo T, Wang Z, Weill JC, Woodgate R (tháng 11 năm 2001). “Eukaryotic DNA polymerases: proposal for a revised nomenclature”. J. Biol. Chem. 276 (47): 43487–90. doi:10.1074/jbc.R100056200. PMID 11579108.Quản lý CS1: nhiều tên: danh sách tác giả (liên kết)
External links
| Wikimedia Commons có thêm hình ảnh và phương tiện truyền tải về DNA polymerase. |

